FishHisto — Histological Image Analysis Platform
Digital platform for automated segmentation and quantification of fish tissue sections for aquaculture and ichthyology research (2021–2023).
Client: IaLINK (Contract Research)
Role: Researcher, Engineer & Developer
Overview
FishHisto is a digital platform for automated histological image analysis in fish biology, developed for aquaculture and ichthyology research applications. The core challenge in histological assessment is that quantifying tissue morphology — fiber lengths, villus heights, structural densities — is currently done manually by trained technicians, making it slow, costly, and operator-dependent. FishHisto replaces this workflow with a computer vision pipeline that segments, measures, and reports key morphological parameters automatically from microscopy images.
Analyzed Tissue Types
The platform handles three main histological domains, each with a dedicated segmentation and measurement pipeline:
Intestinal morphology — Detection and measurement of intestinal villi (mucosal folds), including villus height, crypt depth, and goblet cell density. These parameters are primary indicators of gut health and nutrient absorption efficiency in farmed fish, directly relevant to feed optimization in aquaculture.
Muscle fiber analysis — Segmentation of individual muscle fibers from cross-sectional slides, with automatic measurement of fiber diameter distribution and density. Muscle fiber profiles are key indicators of growth rate and flesh quality.
Foveal structures — Analysis of retinal tissue sections for foveal morphology characterization, relevant to visual ecology and species identification studies.
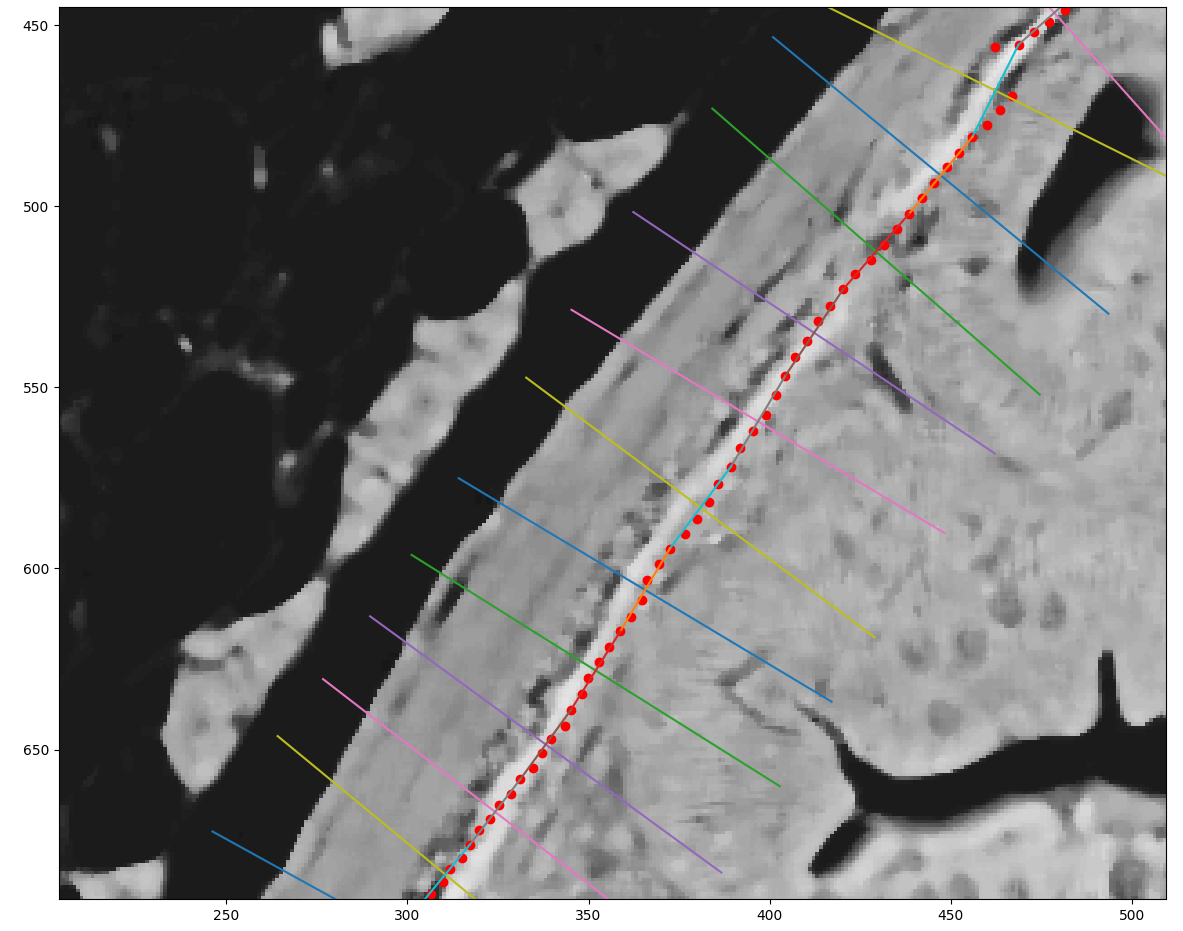
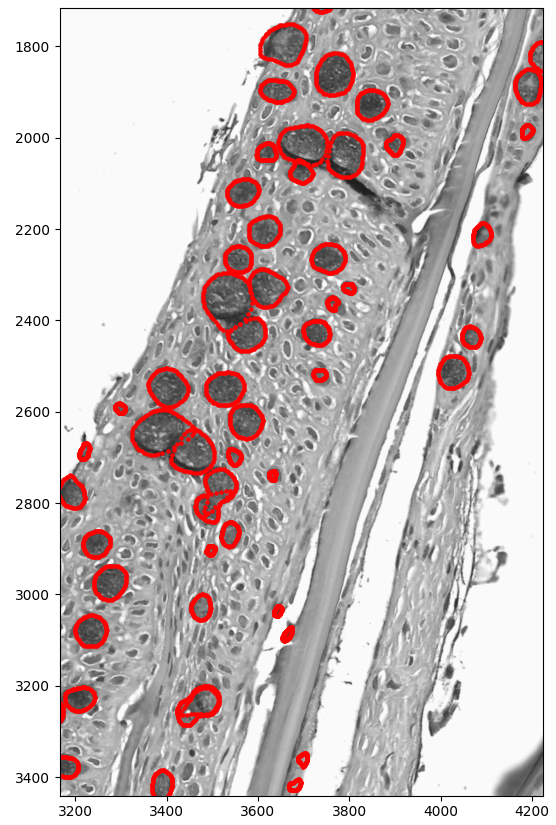
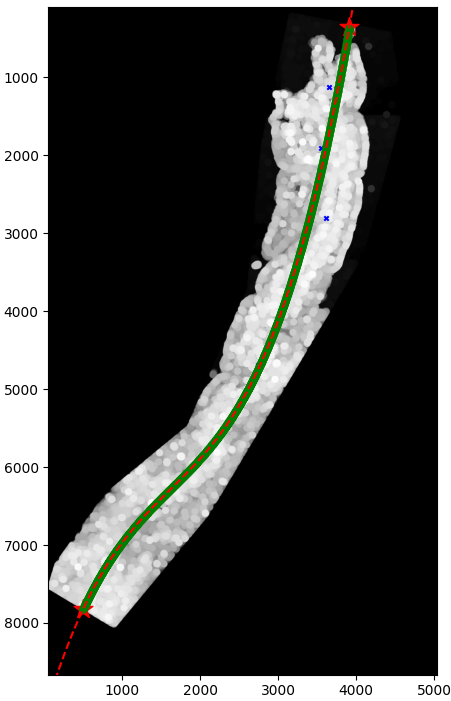

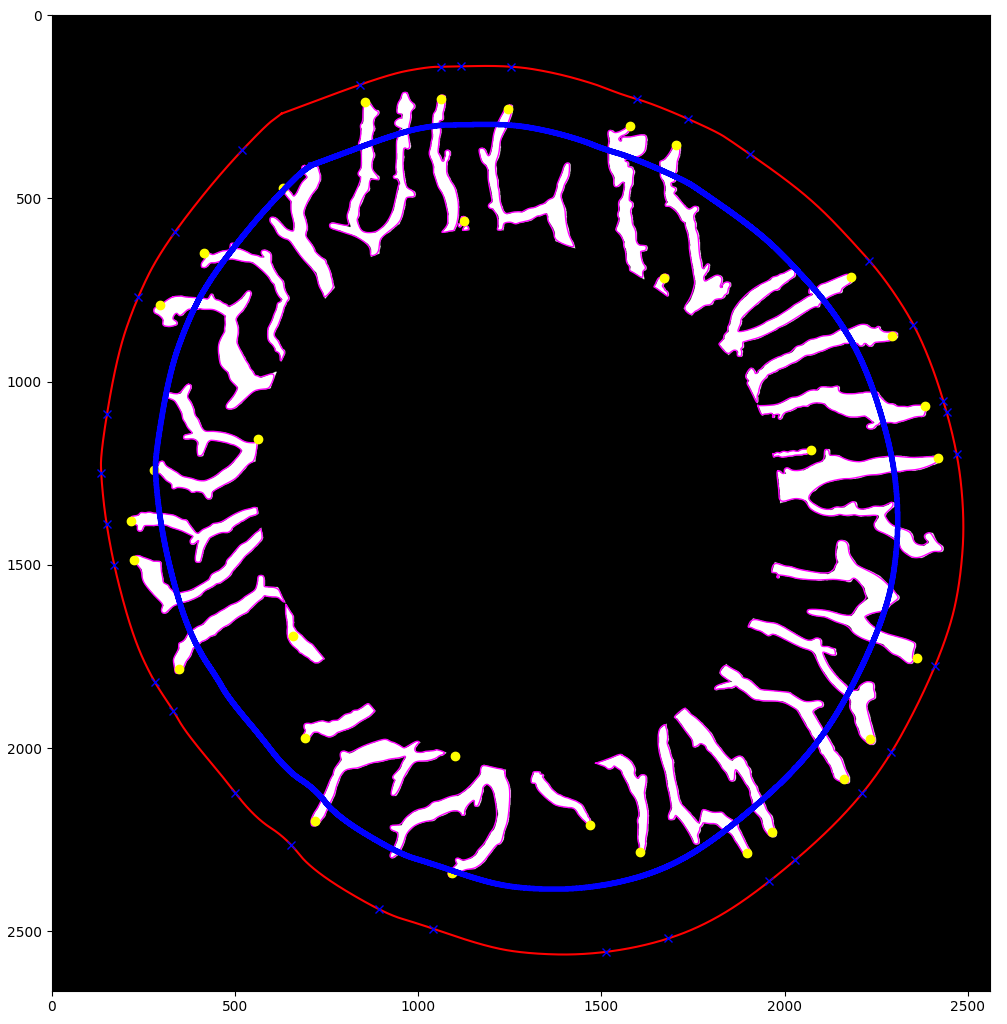
Image Processing Pipeline
The analysis workflow processes whole-slide microscopy images through four sequential stages: preprocessing and stain normalization, tissue boundary detection, structure-level segmentation (individual fibers or villi), and morphometric quantification. Results are exported as structured datasets per slide, enabling batch processing of large experimental cohorts and statistical comparison across treatment groups.