Pediatric e-Health Remote Monitoring Platform
IoT-based remote monitoring system and personalized data mining models for predicting respiratory crisis risk in chronic pediatric patients — FONDEF CONICYT (2014–2016).
Funding: FONDEF Idea — CONICYT, Grant no. CA13i-10300 (2014–2016)
Principal Director: Sebastián A. Ríos, Universidad de Chile
Role: Technical Researcher & Developer
Clinical partner: Hospital Exequiel González Cortés (largest public pediatric hospital in Santiago, Chile)
Motivation
Chronic respiratory diseases in children require continuous monitoring of vital signs to prevent acute deterioration. In Chile, despite broad primary care coverage, remote health assistance technology is practically nonexistent and public hospital overcrowding leaves many patients without timely intervention. This project — known as Proyecto Almohadita — hypothesized that data mining techniques applied to continuous biometric monitoring could anticipate respiratory crisis states in outpatient pediatric patients before a clinical emergency occurs, enabling timely intervention even from home.
IoT Acquisition Platform
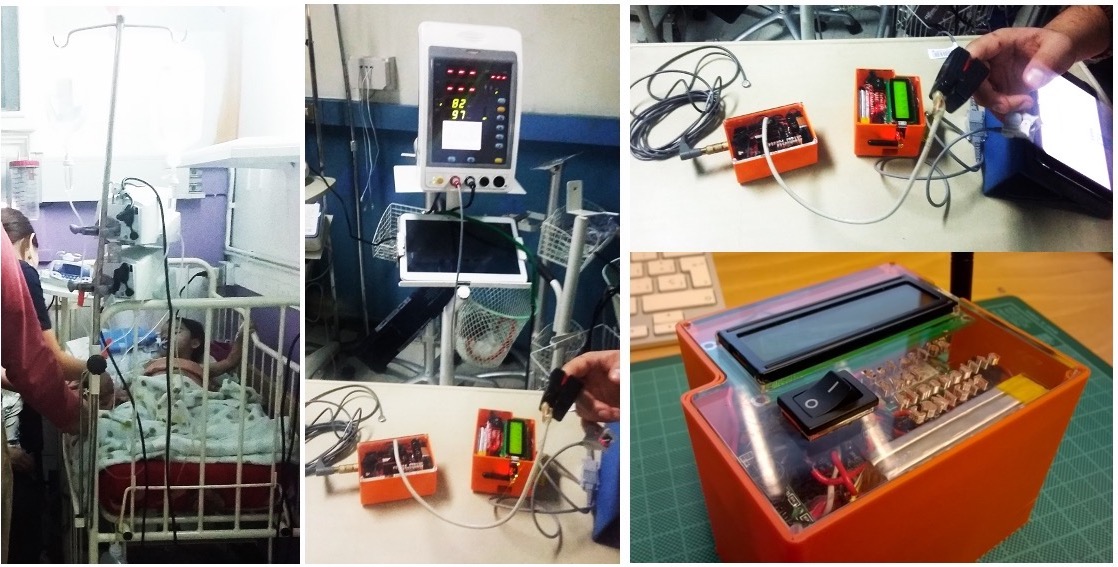
The hardware platform fuses wearable sensors with wireless cloud communication into two complementary devices, built on open-source Arduino hardware.
Hand Remote Unit (HRU) — wearable patient device
Attaches a finger pulse oximeter to an Arduino via an e-Health Sensor Shield, continuously capturing temperature, heart rate (HR), and oxygen saturation (SpO₂). Readings are filtered on-device using a median absolute deviation outlier rejection method (robust to erratic infant movements and sensor misplacement), then packaged and transmitted by radio at 433 MHz at one measurement per second.
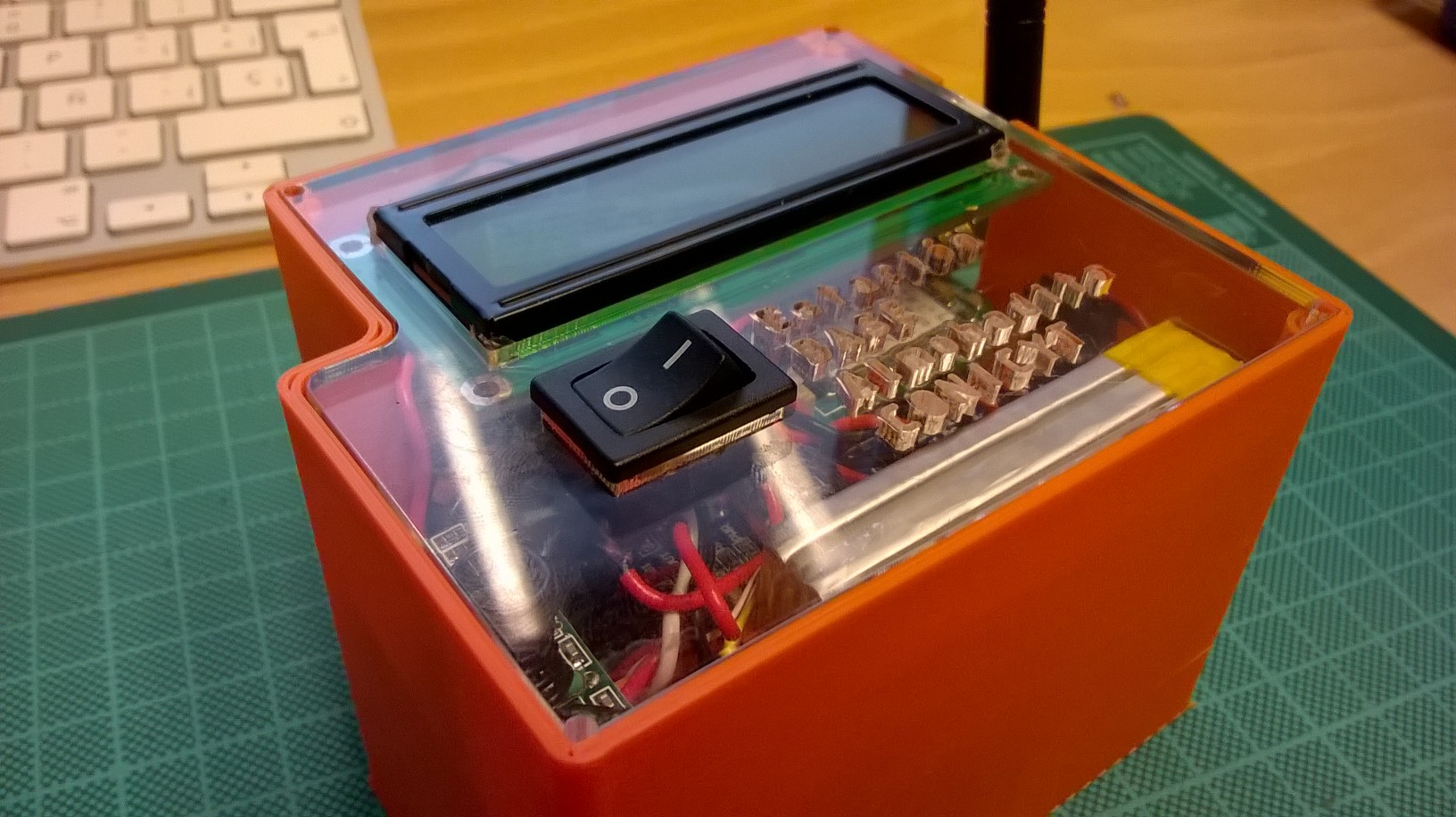
GPRS Central Unit (GPRS-CU) — cloud gateway
A two-Arduino unit receives data from one or more HRUs by radio and manages upload through a GPRS modem. Multiple HRUs can share a single GPRS-CU, reducing ward deployment cost. Once data is uploaded, a server receives it for model prediction.


Predictive Model v2.0
Dataset
Data from 77 anonymized pediatric patients with chronic respiratory diseases, collected from the Pediatrics and Intensive Care units of the hospital. Vital signs — temperature, heart rate (HR), respiratory rate (RR), and oxygen saturation (SpO₂) — were recorded every 2–4 hours depending on the unit. Each record also captures the type of respiratory support, which ranges in severity from:
- Ambient breathing (no support)
- Low-flow nasal cannula
- Halo / Venturi mask
- Mask with recirculation (transitional)
- High-flow nasal cannula (NAF)
- Non-invasive mechanical ventilation (CPAP / BPAP)
- Invasive mechanical ventilation (VMI)
Target Variable: Respiratory Crisis Risk
A key methodological contribution was redefining the risk label. Rather than classifying by current ventilation type, the model labels as “at risk” the vital sign state measured just before an upward transition in respiratory support — i.e., the biometric pattern that actually triggered the clinical team’s escalation decision. This captures the physiological signal that precedes a crisis, not the crisis state itself.
Two-Phase Architecture
Phase 1 — Time Series Forecasting
For each vital sign and each patient, 43 time series models are evaluated across a rolling 3-day window, including Moving Average (MA), Exponential Smoothing, Holt-Winters, Harmonic Seasonal, AR, ARMA, and ARIMA(p,d,q). The model minimizing both mean squared error and maximum deviation is selected per vital sign per patient. This step predicts the patient’s next physiological state.
Phase 2 — Risk Classification
The predicted vital signs are fed into a classifier trained on the same patient’s historical data. Five classifier families are evaluated across all 15 possible input variable combinations (temperature, HR, RR, SpO₂): Naïve Bayes, Artificial Neural Network (ANN), Support Vector Machine (SVM), Logistic Regression, and Decision Tree. The optimal model is selected by maximizing sensitivity, then accuracy, then specificity.
The result is a personalized per-patient pipeline: the optimal time series model and classifier combination differs across patients, reflecting individual physiological patterns — an approach aligned with the emerging global trend toward precision health.
Results
| Metric | Value |
|---|---|
| Patients in dataset | 77 |
| Time series models evaluated per vital sign | 43 |
| Total models evaluated per patient record | 172 |
| Classification models evaluated | 15 per patient |
| Time series prediction error (avg.) | ~10% |
| Classification accuracy (avg.) | ~80% |
These results represent a substantial improvement over the first-year model, which had prediction errors exceeding 30%. The clinical team validated that the model produces no similar prior work in the pediatric chronic disease monitoring literature — both the remote monitoring methodology and the per-patient personalized pipeline are novel contributions.
More information at the Universidad de Chile project page.